Table 1.
Results for E. coli simulation
Click on table to view larger version.
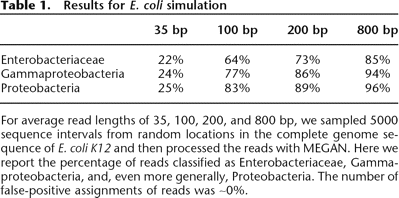
For average read lengths of 35, 100, 200, and 800 bp, we sampled 5000 sequence intervals from random locations in the complete genome sequence of E. coli K12 and then processed the reads with MEGAN. Here we report the percentage of reads classified as Enterobacteriaceae, Gammaproteobacteria, and, even more generally, Proteobacteria. The number of false-positive assignments of reads was ∼0%.











