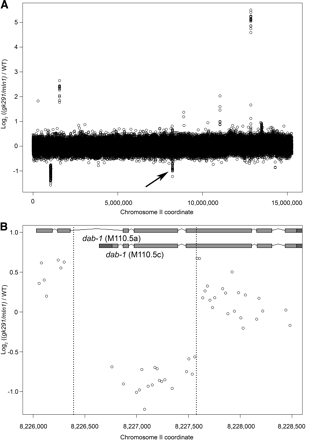
Detection of the 1202-bp deletion in dab-1 (gk291) in a wash-sampled balanced heterozygous population. (A) The normalized log2 ratios [(dab-1(-)/mIn1)/WT] of the average fluorescence intensities for probe pairs to chromosome II are plotted. The arrow indicates the dab-1 deletion (other features are discussed in the text). (B) Normalized fluorescence ratios for probe pairs targeting dab-1 are shown. Sequenced deletion breakpoints are indicated by dotted lines and were accurately predicted by CGH. The left breakpoint lies within the second intron. Overlapping probes targeting the right breakpoint span just 73 bp, allowing resolution of the right breakpoint to within fewer than 50 bp.











