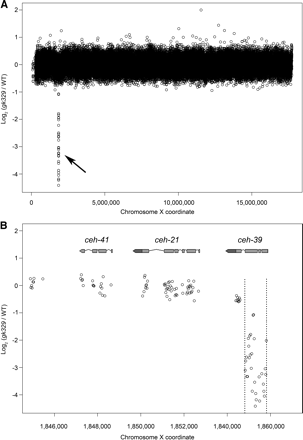
Detection of a 1047-bp homozygous viable deletion in ceh-39 (gk329). (A) Normalized log2 ratios (gk329/WT) for the average fluorescence intensities for all probe pairs to the X chromosome are shown. The arrow indicates the deletion. (B) Intensity ratios for probes to ceh-39, ceh-21, and ceh-41 are shown with WormBase gene models to illustrate probe coverage in exons near the deletion. Sequenced deletion breakpoints are indicated by dotted lines. CGH accurately identified the left breakpoint between exons 3 and 4 of ceh-39.











