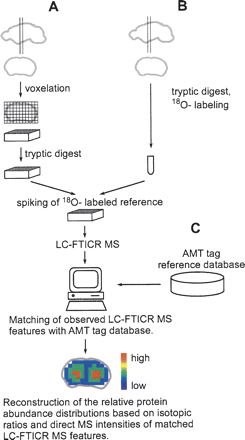
Strategy for spatial mapping of protein abundance patterns in the mouse brain using an example of a coronal slice. (A) Coronal section was further dissected into voxels (1-mm3 cubes) followed by tryptic digestion of each voxel sample. (B) The entire coronal slice was digested by trypsin followed by 18O-labeling and spiking into each voxel sample as a reference for robust quantitation. (C) The observed LC-FTICR features were matched against an AMT tag database containing theoretical masses and observed elution times of previously identified peptides from LC-MS/MS analyses.











