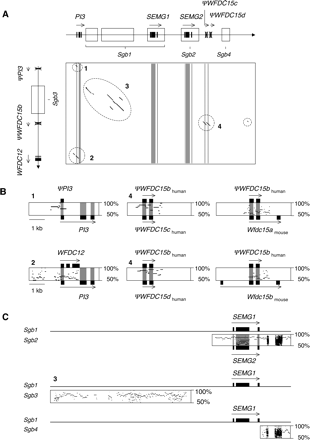
Sequence conservation between paralogous segments in the centromeric WFDC sublocus. (A) A dot plot depicts sequence conservation between the two paralogous segments that reside within the human centromeric WFDC sublocus. The X- and Y-axes represent the PI3-to-ΨWFCD15d and WFDC12-to-ΨPI3 duplicons, respectively (boxed regions in Fig. 1B). A schematic representation of the exon-based gene structures and pseudogenes is shown for each, with the positions of exons along the X-axis highlighted in the dot plot. The four SEMG genomic blocks (Sgb) are boxed and labeled. Sgb1 is larger than previously described (Ulvsback et al. 1992) and is fragmented into three pieces, as determined by comparison with the lemur sequence. The two paralogous segments have significant coding and noncoding sequence conservation (highlighted by dashed circles, four of which are labeled 1–4 and detailed in B and C). (B) PipMaker plots of three paralogous regions highlighted in A. (Left column, panel 1) ΨPI3 resides within a genomic segment that is 79% similar to a 774-bp region of PI3, including exon 1; (left column, panel 2) there is considerable conservation between PI3 and WFDC12, including the proximal 1495 bp of the 5′region (58% identical), the first exon (79% identical at the nucleotide level; 10/22 residues identical at the amino acid level), and the first PI3 intron with WFDC12 exons 2 and 3 (55% identical); (center column, panels 4) the three human WFDC15 pseudogenes show >79% sequence identity with each other as well as lower but significant identity with a 1.2-kb segment of mouse Wfdc15a and Wfdc15b (right column). (C) PipMaker plots showing the sequence conservation among the four paralogous human SEMG genomic blocks (Sgb1 to Sgb4). In all cases, the reference sequence is Sgb1. Newly identified Sgb3 and Sgb4 are truncated blocks that have lost all SEMG-coding sequences. The Sgb1/Sgb3 plot corresponds to the highlighted region 3 in panel A. Panels A and C are not to scale.











