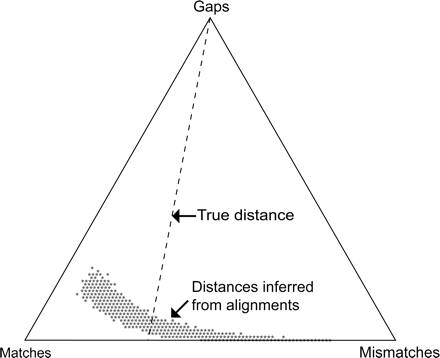
A barycentric representation of the distribution of matches, mismatches, and gaps for optimally aligned pairs of sequences having exactly 60 matches, 30 mismatches, and 10 gaps each over the entire range of alignment parameters (match, mismatch, and gap penalties) for which mismatches are penalized more than matches. Each point in the triangle represents a number of matches, mismatches, and gaps summing to 100. The lengths of the perpendiculars from a point in the diagram to the right, left, and bottom side of the triangle are proportional to the corresponding match, mismatch, and gap fractions, respectively. Simulation outcomes are represented by the dark gray points near the bottom of the triangle. In 1000 trials, the true sequence-pair description (60 matches, 30 mismatches, 10 gaps) was never recovered, and only those 1.1% of the alignments lying precisely on the line labeled “true distance” yielded estimated distances equal to the true distance (1/3) between the original pair of sequences. Adapted, with permission, from Figure 2.3 of Fleissner 2003.











