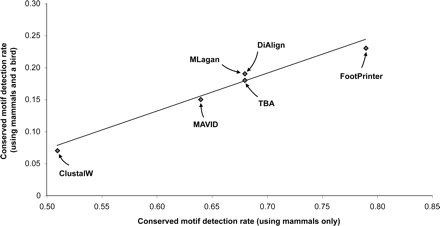
Graph showing the performance of different multiple sequence alignment programs in detecting conserved motifs of length 10 in sets of orthologous 1000 base-pair putative promoter regions of vertebrate genomes (Prakash and Tompa 2005). The axes represent the fraction of alignments (using the given program) of orthologous sequences in which at least one perfectly conserved motif of length 10 was detected. The x-axis represents 5073 alignments containing human, chimp, mouse, and rat sequences, while the y-axis represents 945 alignments of human, chimp, mouse, rat, and chicken. The smaller number of hits in the latter set is partly attributable to fewer conserved motifs and partly to increased difficulty of detection. The programs used are ClustalW (Thompson et al. 1994), MAVID (Bray and Pachter 2004), DiAlign (Morgenstern 1999), MLagan (Brudno et al. 2003a), TBA (Blanchette et al. 2004), and FootPrinter (Blanchette and Tompa 2003). Data are from Figure 2 of Prakash and Tompa (2005).











