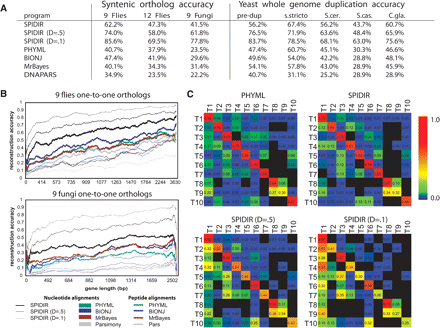
SPIDIR learning methodology leads to significantly higher accuracy. (A) Comparison of SPIDIR and several popular phylogenetic methods for syntenic orthologs in nine flies, 12 flies, and nine fungi, and for duplicate genes arising from whole-genome duplication. “pre-dup” gives the accuracy of reconstructing the topology of the three preduplicated species, “s. stricto” is the topology accuracy of only the four sensu strict species, and “S. cer”, “S. cas”, and “C. gla” give the accuracy of placing annotated ohnologs of each species on opposing sides of the whole-genome duplication node. (B) Reconstruction accuracy for SPIDIR correlates with gene length, similarly to other methods, and is consistently higher. (C) Reconstruction accuracy for SPIDIR and PHYML for ROSE simulated fly alignments according to 10 most frequent topologies T1–T10 (left). Both methods are unbiased and recover alternate topologies at similar rates (top), although SPIDIR was trained on T1. With increasing duplication cost, T1 becomes favored by SPIDIR (D = 0.5 and 0.1).











