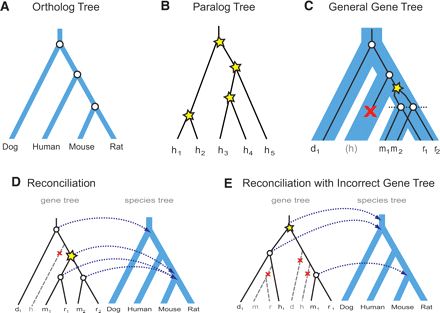
Relationship between gene trees and species trees. (A) Ortholog trees used to study species evolution. Each internal node represents a speciation event (circle). (B) Paralog trees used to study gene family expansions within a single species. Each internal node represents a duplication event (star). (C) General gene trees combine both orthologs and paralogs across multiple species to infer gene duplication (star), gene loss (×), and speciation (circle) events. Each gene is named with the first letter of the corresponding species. The gene tree (black lines) can be viewed as evolving inside the species tree (blue area), implying coordinated speciation events at branching points in the species tree (dotted line). (D) Gene duplication and loss events are inferred by reconciling a gene tree to a species tree, mapping each gene-tree node to its closest species-tree common ancestor node (arrows). (E) When the gene tree is incorrect, many spurious events will be inferred. In this example, a common misplacement of rodents due to long-branch-attraction leads to four spurious events (one duplication and at least three losses).











