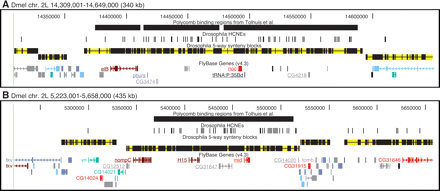
HCNE-clusters spanning coregulated genes and boundary agreement among synteny blocks, HCNE clusters, and Polycomb binding regions. Gene models are colored by predicted core promoter type as in Fig. 2. Only selected genes are labeled. (A) The paralogous zinc finger genes elB and noc, implicated in tracheal and appendage development, have different, but partially overlapping, spatial expression patterns during embryonic development (Dorfman et al. 2002) and are coexpressed in larval leg and wing discs (Weihe et al. 2004). Among the five flies, elB and noc are in conserved microsynteny with a tRNA gene and at least three protein-coding genes (underlined), which have no evidence of being functionally related to elB or noc: pburs encodes a subunit of the hormone bursicon required for wing expansion and associated cuticle changes after flies emerge from pupae (Luo et al. 2005); CG3474 is predicted to encode a cuticle component; CG4218 is predicted to encode a protein of unknown function. (B) The paralogous T-box genes H15 and mid are involved in regulation of heart development and have similar spatial expression patterns during embryonic development (Miskolczi-McCallum et al. 2005; Reim et al. 2005). They are in conserved microsynteny with four other genes (underlined) among the five flies: CG12512, predicted to encode a protein with an AMP-dependent synthetase and ligase domain; nompC, encoding a mechanosensory transduction channel (Walker et al. 2000); and two genes of unknown function. The developmental regulators vri (George and Terracol 1997) and tomb (Jiang et al. 2007) are centrally positioned in neighboring synteny blocks. Two transcript isoforms are shown for tkv because it has two major transcription start sites with different types of core promoter predictions.











