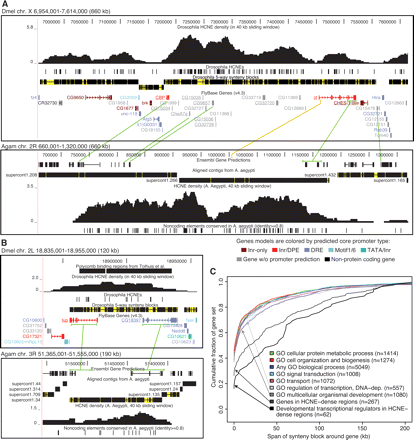
Genes associated with HCNE arrays tend to be in large fly-mosquito synteny blocks. (A,B) Examples of synteny blocks. Gaps within synteny blocks are colored yellow. Green lines connect genes in conserved microsynteny between Dmel and Agam. Microsynteny conservation between Dmel and Agam was determined by examining chained BLASTZ and TBLASTN alignments in the UCSC Genome Browser (Kent et al. 2002). Sometimes only parts of genes could be matched (e.g., in the case of ct). Aaeg contigs aligned to the Agam assembly are shown with regions having ≥50% identity over 50 bp in black and other regions in yellow. HCNE densities were computed as the fraction of bases in HCNEs in sliding windows of 40 kb. The UCSC Genome Browser (Kent et al. 2002) was used in making the images. (A) The ct locus in Dmel (upper panel) and Agam (lower panel). ct and nine other genes (underlined) show strong evidence of being in conserved microsynteny among the five flies. The orange line indicates a noncoding BLASTZ match between Dmel and Agam and hints at the location of the first ct exon in Agam. Comparison of HCNE density curves also supports that the first ct exon in Agam is in the area indicated by the orange line. Supporting a common origin of the HCNE clusters at the ct loci in flies and mosquitoes, the HCNE density curves have similar shapes. Two density peaks are visible in both organisms: one between CG9657 and ct, and the other within the borders of ct. The developmental transcriptional regulator brk (Moser and Campbell 2005) is centrally positioned in an adjacent synteny block. CG9650, which dominates a neighboring HCNE-rich synteny block, is expressed in developing CNS and PNS and encodes a putative C2H2 zinc finger protein (McGovern et al. 2003). (B) The tailup (tup) locus in Dmel (upper panel) and Agam (lower panel). tup is in conserved microsynteny with CG18397 among the five flies, Agam and Aaeg. tup encodes a homeodomain transcription factor involved in development (Thor and Thomas 1997). CG18397 is predicted to encode a protein with an AMP-dependent synthetase and ligase domain. In both flies and mosquitoes, HCNEs are found throughout the synteny block. Some HCNEs are within introns of tup and CG18397. This, combined with the lack of evidence for a functional relationship between the two genes, indicates that they have been kept in proximity in order to maintain the HCNE array. (C) For each gene that we could assign to a synteny block, we measured the span of its synteny block excluding the region spanned by the gene itself (in order to control for differences in gene size). Each curve shows the cumulative distribution of synteny block span, measured in Dmel bp, around genes in a particular category. Categories were defined from GO biological process annotation and HCNE density. The category “any biological process” contains all genes annotated with a GO biological process term other than “biological process unknown.” Genes in HCNE-dense regions overlap a 40-kb region where at least 1% (400 bp) of the sequence is in HCNEs. Numbers within parentheses indicate the number of genes annotated to the indicated process and assigned to a single synteny block.











