Categorization of existing gene annotations according to comparative evidence
Click on table to view larger version.
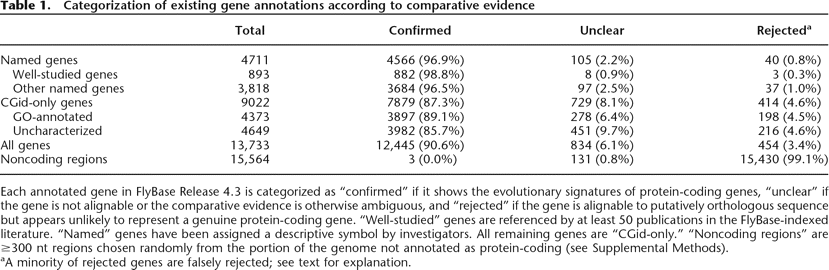
Each annotated gene in FlyBase Release 4.3 is categorized as “confirmed” if it shows the evolutionary signatures of protein-coding genes, “unclear” if the gene is not alignable or the comparative evidence is otherwise ambiguous, and “rejected” if the gene is alignable to putatively orthologous sequence but appears unlikely to represent a genuine protein-coding gene. “Well-studied” genes are referenced by at least 50 publications in the FlyBase-indexed literature. “Named” genes have been assigned a descriptive symbol by investigators. All remaining genes are “CGid-only.” “Noncoding regions” are ≥300 nt regions chosen randomly from the portion of the genome not annotated as protein-coding (see Supplemental Methods).
aA minority of rejected genes are falsely rejected; see text for explanation.











