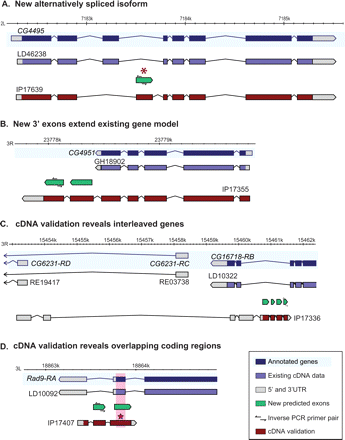
Full-length cDNA sequences recovered from exon predictions through inverse PCR. (A) Alternatively spliced transcripts—Exon Shuffling. The clone, IP17639, validates prediction congochr2L7183503 and provides evidence for an alternative transcript of the gene CG4495. Analysis of the embryonic microarray data (Manak et al. 2006) shows this exon is not used in embryogenesis, suggesting stage-specific splicing. Interestingly, the two alternative exons encode 20 identical amino acids at the N-terminal side of the exon. (B) 3′ CDS extension. The clone, IP17355, validates two predicted exons, congochr3R23777966 and congochr3R23778197, and provides evidence for an alternative transcript encoding an additional 126 aa at the C-terminal end of the gene, CG4951. In addition, the clone contains 185 bp of 3′ UTR. (C) New spliced interleaved gene. The clone, IP17336, validates four predicted exons, congochr3R15461397, congochr3R15461180, congochr3R15461031, and congochr3R15460742, and provides evidence for four additional exons. (D) Novel spliced overlapping gene. The clone, IP17407, validates prediction congochr3L18835687 and extends the CDS by 22 aa at the N terminus and 79 aa at the C terminus. The third coding exon overlaps the coding sequence of the gene on the opposite strand, Rad9, such that 45 aa on each strand are encoded in the region of overlap.











