Start/stop codon drift for genes with big and small initial protein-coding exons, with and without CpG islands, and with and without alternative promoters
Click on table to view larger version.
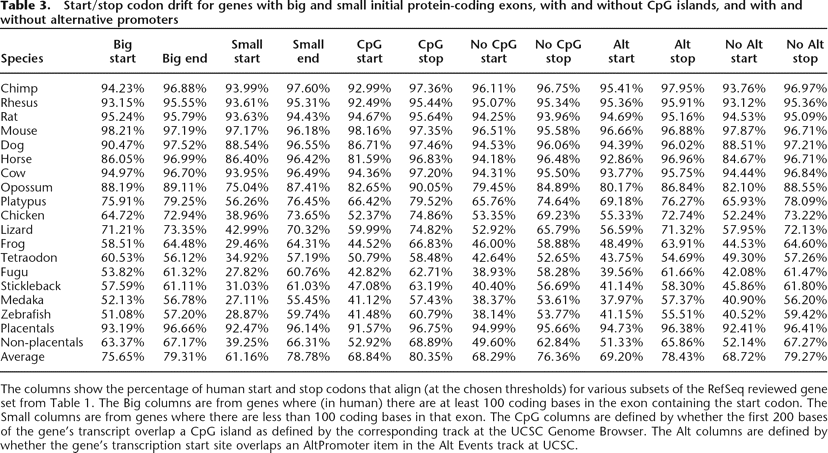
The columns show the percentage of human start and stop codons that align (at the chosen thresholds) for various subsets of the RefSeq reviewed gene set from Table 1. The Big columns are from genes where (in human) there are at least 100 coding bases in the exon containing the start codon. The Small columns are from genes where there are less than 100 coding bases in that exon. The CpG columns are defined by whether the first 200 bases of the gene’s transcript overlap a CpG island as defined by the corresponding track at the UCSC Genome Browser. The Alt columns are defined by whether the gene’s transcription start site overlaps an AltPromoter item in the Alt Events track at UCSC.











