Species-by-species information
Click on table to view larger version.
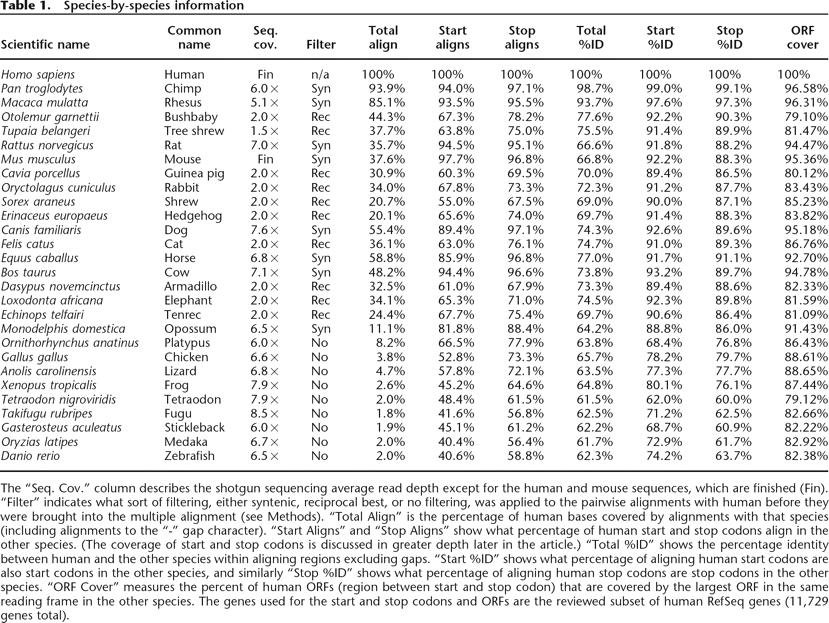
The “Seq. Cov.” column describes the shotgun sequencing average read depth except for the human and mouse sequences, which are finished (Fin). “Filter” indicates what sort of filtering, either syntenic, reciprocal best, or no filtering, was applied to the pairwise alignments with human before they were brought into the multiple alignment (see Methods). “Total Align” is the percentage of human bases covered by alignments with that species (including alignments to the “-” gap character). “Start Aligns” and “Stop Aligns” show what percentage of human start and stop codons align in the other species. (The coverage of start and stop codons is discussed in greater depth later in the article.) “Total %ID” shows the percentage identity between human and the other species within aligning regions excluding gaps. “Start %ID” shows what percentage of aligning human start codons are also start codons in the other species, and similarly “Stop %ID” shows what percentage of aligning human stop codons are stop codons in the other species. “ORF Cover” measures the percent of human ORFs (region between start and stop codon) that are covered by the largest ORF in the same reading frame in the other species. The genes used for the start and stop codons and ORFs are the reviewed subset of human RefSeq genes (11,729 genes total).











