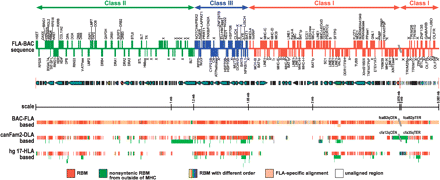
FLA annotation from finished BAC sequence (RPCI 86 BAC library constructed from the DNA of a domestic cat, Gus) and 1.9× WGS contigs alignment with three MHC models (Yuhki et al. 2003; Beck et al. 2005). Two segments of cat MHC (FLA) sequences—2.976-Mb B2cen MHC sequence and 0.0362-Mb B2ter MHC sequence—were assembled based on the human HLA sequence (The MHC Sequencing Consortium 1999). Gene annotation of this FLA sequence was performed using the GENSCAN (Burge and Karlin 1997) program. Genes with forward and reverse orientations were placed as solid blocks above and below the line, respectively. A gap position of these two FLA sequences is indicated as double slash lines. Contigs from the 1.9× WGS genome sequence were aligned using the Cross_match program with three models—FLA, DLA (canFam2), and HLA (hg17 assembly), respectively. These three sets of contig alignments were compared and highlighted with different colors with the following categories: (red) conserved sequence blocks with the same order between FLA and DLA shown on FLA and DLA lines or between FLA and HLA shown on the HLA line; (green) nonconserved sequences between FLA/DLA or FLA/HLA but a part of FLA sequences conserved in other genomic regions in dog and human and translocated to MHC regions in DLA or HLA; (multiple colors) conserved sequence blocks between FLA/DLA or FLA/HLA with different order; (pink) FLA-specific blocks; (white) no 2× WGS contigs were aligned in DLA or HLA.











