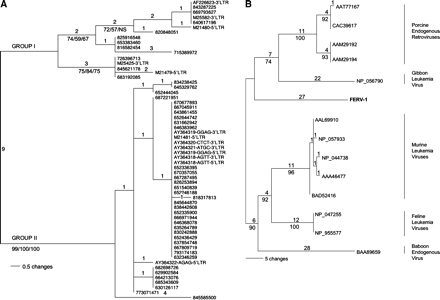
Phylogenetic relationship among endogenous retroviruses. (A) Endogenous FeLV present in the genome of the domestic cat. Analysis is based on a 274-bp alignment of 13 previously published proviral LTRs (GenBank accessions) and 47 cat genomic sequences (trace IDs). Strong bootstrap support suggests that the proviral sequences fall into two groups, as labeled. The maximum parsimony tree is shown (length = 39, CI = 0.923, RC = 0.911). Bootstrap support (>70%) is indicated for maximum parsimony, neighbor joining, and maximum likelihood methods. (B) FERV-1. Phylogenetic relationships among previously identified retroviruses and a novel retroelement sequence in the cat genome. One copy of the novel provirus (FERV-1) in an unpublished BAC sequence was used to generate the maximum parsimony (MP) tree (length = 142, CI = 0.915, RC = 0.851), based on the predicted sequence of 205 amino acids of Pol. The number of steps is listed above the branches, while MP bootstrap support is indicated below the branches of the major clades.











