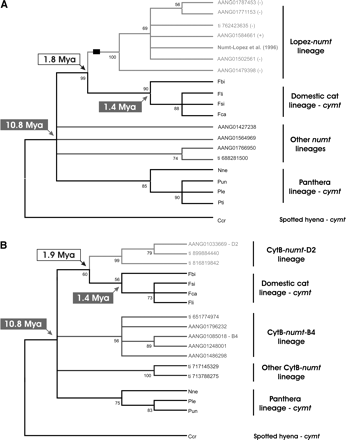
Phylogenetic analyses of felid mitochondrial (cymt) and homologous domestic cat numt sequences. Divergence dates of numt lineages-defining nodes were estimated following Lopez et al. (1994). Divergence dates (in gray boxes) assumed for the Felidae radiation and the represented species of the domestic cat lineage were retrieved from Johnson et al. (2006). Bootstrap values, calculated from 1000 replications, are placed at each branchpoint. (A) Minimum-evolution condensed tree (50% cutoff value) of a 570-bp segment of the mtDNA NADH dehydrogenase subunit I (NDI is a gene segment represented in the Lopez-numt) and homologous numt detected by BLAST searches in F. catus. (Black rectangle) The phylogenetically informative 10-bp deletion shared by all representatives of the Lopez-numt lineage. The symbols (+) and (−) represent the orientation of the genomic DNA sequence. The numt sequences are labeled using trace (ti), scaffold, GenBank accession, and chromosome. (B) Minimum-evolution condensed tree (50% cutoff value) of a 426-bp segment of the mtDNA cytochrome b (CytB is a gene segment not represented in the Lopez-numt) and several homologous numts detected by BLAST searches in F. catus. The cymt sequences are labeled with a three-letter code: (Fca) F. catus, domestic cat; (Fsi) Felis silvestris, European wild cat; (Fli) Felis libyca, African wild cat; (Fbi) Felis bieti, Chinese desert cat; (Nne) Neofelis nebulosa, clouded leopard; (Pti) Panthera tigris, tiger; (Pun) Panthera uncia, snow leopard; (Ple) Panthera leo, lion; and outgroup (Ccr) Crocuta crocuta, spotted hyena.











