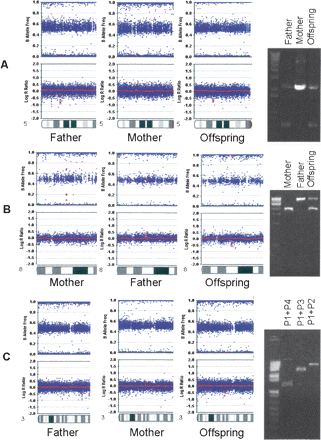
(A) A predicted ∼700-bp CNV within an intronic region of the FBXL7 gene; (B) a predicted ∼1-kb CNV within an intronic region of the EYA1 gene; and (C) a predicted ∼4-kb CNV within an intronic region of the CTDSPL gene are inherited from parent to offspring. The scatterplots for log R Ratio and B Allele Frequency are shown for the father, mother, and offspring; (red dots) the SNPs within the CNVs. The presence of CNVs and their copy numbers are validated by PCR amplification of the region encompassing breakpoints for FBXL7 and EYA1, or by PCR primer walking for CTDSPL (see Fig. 4 for more detail on primer locations).











