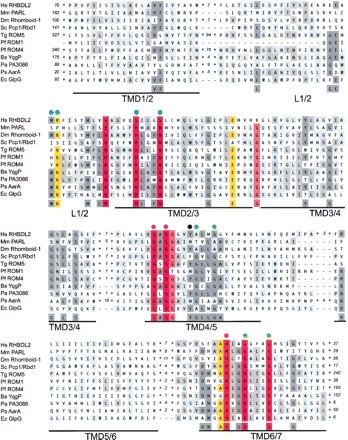
Functionally based rhomboid protease consensus. Conserved membrane integral portion of rhomboids that have been used in mutagenesis experiments (Urban et al. 2001, 2002b; Esser et al. 2002; McQuibban et al. 2003; Brossier et al. 2005; Lemberg et al. 2005; Maegawa et al. 2005; Urban and Wolfe 2005; Baker et al. 2006; Cipolat et al. 2006; O’Donnell et al. 2006; Stevenson et al. 2007) were aligned. For human (Homo sapiens, Hs) RHBDL2, Drosophila melanogaster (Dm) Rhomboid-1, Toxoplasma gondii (Tg) ROM5, Plasmodium falciparum (Pf) ROM1 and ROM4, Escherichia coli (Ec) GlpG, Providencia stuartii (Ps) AarA, Pseudomonas aeruginosa (Pa) rhomboid PA3086, and Bacillus subtilis (Bs) YqgP TMD1 to TMD6 are shown; for mouse (Mus musculus, Mm) PARL and Saccharomyces cerevisiae (Sc) Pcp1/Rbd1, the topologically equivalent TMD2 to TMD7 were aligned. For accession numbers see below. Alignment was performed by ClustalW and corrected manually to remove gaps in predicted TMDs and loops extending the sequence of the E. coli GlpG rhomboid. The more variable N- and C-terminal domains (including additional TMDs) were excluded from the alignment (number of amino acids is indicated). TMD predictions were based on the structure of the E. coli rhomboid GlpG and are underlined (Wang et al. 2006). The L1 loop (L2 for PARL-type rhomboids) containing widely conserved and functional important residues is indicated; invariant residues forming the active site (GxSx and H) are labeled by red dots; residues affecting the activity only in certain rhomboids or experimental conditions are highlighted by blue dots; position of a tyrosine residue of the E. coli GlpG rhomboid (Y205) suggested to be involved in positioning of the catalytic histidine (H254) (Wang et al. 2006) is labeled by a black dot; small residues such as glycines (G) and alanines (A) in TMD4/6 and TMD6/7 that allow tight helix packing are labeled by green dots (Ben-Shem et al. 2007). The background color reflects the degree of identity/similarity of sequence alignment (100%, red; 90%–99%, light-red; 80%–89%, yellow; 50%–79%, dark gray; 30%–49%, light gray). For accession numbers for the eukaryotic rhomboids, see Figure 3 and Table S1. The accession number for E. coli GlpG is Swiss-Prot:P09391; P. stuartii AarA is Swiss-Prot:P46116; P. aeruginosa PA3086 is Swiss-Prot:Q9HZC2; and B. subtilis YqgP is Swiss-Prot:P54493.











