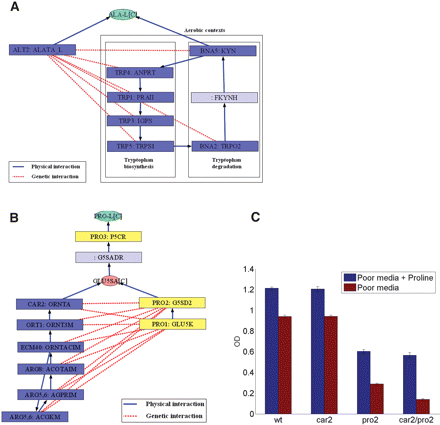
Functional pathways of alanine and proline biosynthesis as reflected by CDA. (A) A functional pathway of alanine biosynthesis under aerobic and anaerobic poor media. Rectangular nodes represent metabolic reactions, specifying names of the coding genes and names of the reactions. The circular node represents the metabolite alanine. Blue edges represent physical interactions between enzymes, in the form of a metabolite that is the product of one enzyme and the substrate of the other. Red edges represent genetic interactions between genes that are specific to alanine biosynthesis. (B) A functional pathway of proline biosynthesis in aerobic minimal media condition. (C) Proline auxotrophy experiment for the CAR2 and PRO2 single deletion, and CAR2/PRO2 double deletion strains. The experiments show a significant drop in maximum optical density (OD) for the CAR2/PRO2 double mutant strain when proline is removed from the growth medium. The results show the existence of a synthetic sick interaction between CAR2 and PRO2 in minimal media that lacks proline.











