Summary of information measures for six tagsets in SeattleSNPs/PARC samples
Click on table to view larger version.
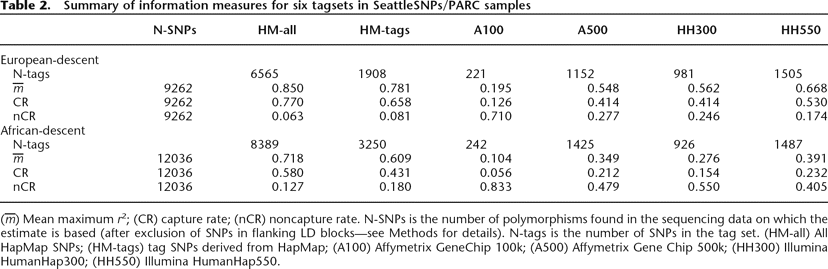
(m̅) Mean maximum r2; (CR) capture rate; (nCR) noncapture rate. N-SNPs is the number of polymorphisms found in the sequencing data on which the estimate is based (after exclusion of SNPs in flanking LD blocks—see Methods for details). N-tags is the number of SNPs in the tag set. (HM-all) All HapMap SNPs; (HM-tags) tag SNPs derived from HapMap; (A100) Affymetrix GeneChip 100k; (A500) Affymetrix Gene Chip 500k; (HH300) Illumina HumanHap300; (HH550) Illumina HumanHap550.











