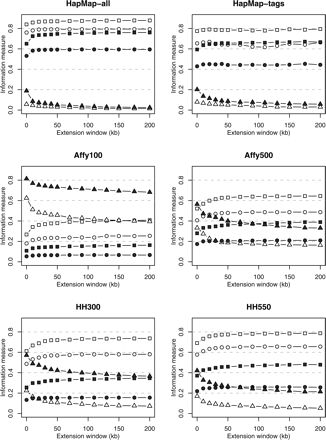
Figure 1.
Information measures for 78 genes using combined SeattleSNPs/PARC and HapMap data for extension windows of 0–200 kb around the sequenced region. Mean maximum r2, capture rates, and noncapture rates are shown by squares, circles, and triangles, respectively; European and African descent samples are distinguished by solid and open symbols, respectively.











