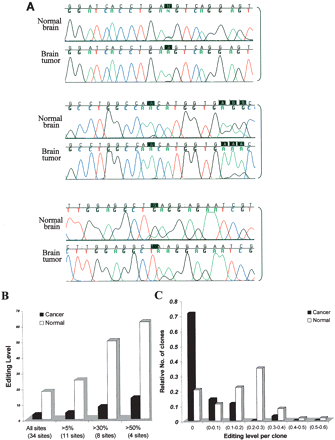
Reduced editing levels at Alu sequences in a human brain tumor. (A) Three representative clusters from the direct sequencing of an Alu sequence located in intron 9 of the MED13 gene of normal and tumor brain (low-grade glioma) tissues are illustrated. Black boxes indicate the editing sites. (B) Alu editing in individually cloned transcripts from normal and malignant brain tissues. PCR products derived from the normal and astrocytoma samples (shown in panel A) were cloned and sequenced. One-hundred-twenty-one clones were sequenced (46 clones from astrocytoma and 75 clones from normal brain tissue). Thirty-four “As” were shown to be edited at least in one of the clones. The percent of editing was analyzed in all sites and is displayed according to cutoff values of >5%, 30%, and 50% editing. White bars represent transcripts from normal brain tissue, and black bars represent clones from brain tumor. (C) Analysis of editing level of Alu sequences in intron 9 of MED13 in individual clones. MED13 transcripts from both normal and malignant brain samples were cloned and sequenced (same samples as in panel A); 71% of the cancer clones, compared to only 21% of the normal clones, do not undergo editing at any site.











