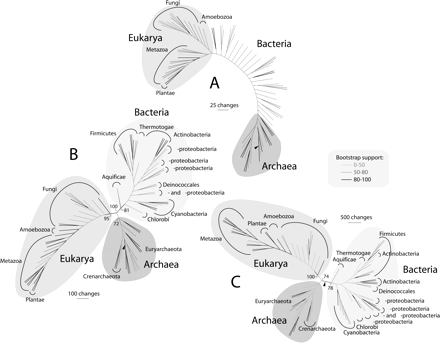
Optimal most-parsimonious phylogenomic trees of proteomes from 82 free-living organisms, generated using subsets of FSF corresponding to different phases of evolutionary history. (A) Ancient FSF, ndFSF < 0.174 (6727 steps; CI = 0.232, RI = 0.687; g1 = −0.316). (B) Intermediate FSF, 0.174 < ndFSF < 0.489 (38,405 steps; CI = 0.184, RI = 0.681; g1 = −0.299). (C) Young FSF, ndFSF > 0.489 (67,555 steps; CI = 0.234, RI = 0.709; g1 = −0.576). Terminal leaves are not labeled, as they would not be legible. Individual trees with taxon labels are shown in Supplemental Figure S3. Bootstrap support (BS) levels for branches are indicated with different shades and with numbers in nodes delimiting superkingdoms.











