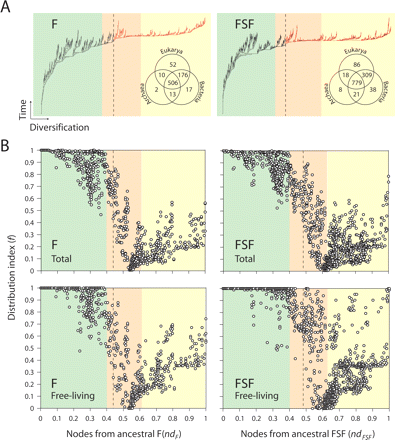
Architectural chronologies of (F) folds (left) and (FSF) fold superfamilies (right) suggest three evolutionary epochs in the timeline of the protein world. (A) Optimal (P < 0.01) most-parsimonious F (85,644 steps; CI = 0.043, RI = 0.770; g1 = −0.134) and FSF (118,119 steps; CI = 0.031, RI = 0.759; g1 = −0.099) trees were reconstructed from a protein domain census in 185 completely sequenced genomes. Venn diagrams show occurrence of architectures in the three superkingdoms of life, Archaea (A), Bacteria (B), and Eukarya (E). Terminal leaves were not labeled, as they would not be legible. (Red) Branches defining F and FSF that occur after the appearance of the first architecture unique to a superkingdom (B). (B) Distribution index of individual architectures (f, the number of species using an architecture/total number of species) against the age of architectures (nd, number of nodes from the root/total number of nodes in the tree) uncovers evolutionary patterns of architectural innovation and usage when studying all genomes or only those that are free-living. Based on these patterns, we propose three evolutionary epochs of the protein world: (light green) structural diversification; (salmon) superkingdom specification; (yellow) organismal diversification epochs.











