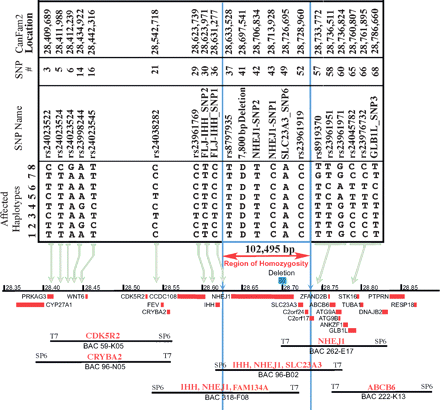
Alignment of SNP haplotypes and a canine BAC contig to the CanFam2 genomic sequence for the cea interval on canine chromosome 37. Eight haplotypes, spanning >376 kb on CFA37, represent cea-transmitting chromosomes segregating in four different breeds and define a linkage disequilibrium interval common to all known cea-affected breeds. Haplotype 1 was observed in all cea-affected Rough and Smooth Collies and Australian Shepherds tested, and in some but not all Border Collies. Haplotypes 2 through 7 were identified in specific Border Collies, and haplotype 8 was only seen in European Shetland Sheepdogs. All haplotypes are identical, and all affected dogs are homozygous for SNPs rs8797935 through rs23961919 (the region bounded by vertical blue lines). (Blue box above the chromosome, the black line in the center of the figure) The position of the 7799-bp deletion in intron 4 of NHEJ1. (Red boxes below the chromosome line) Genes in the region. BACs are tiled below the genes and are identified by their number. The gene-specific probes used to pick the BACs and align them to the chromosome are listed in red above the BAC.











