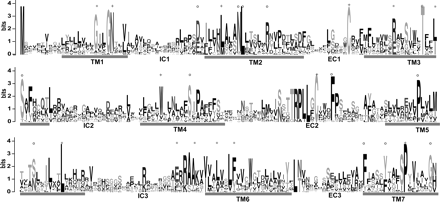
Conserved sequence motifs of the Ora family. Conservation of predicted amino acid sequence for the fish Ora repertoire is displayed as a sequence logo. In this representation, the relative frequency with which an amino acid appears at a given position is reflected by the height of its one-letter amino acid code in the logo, with the total height at a given position proportional to the level of sequence conservation. The regions corresponding to the transmembrane (TM) domains and the intracellular and intracellular domains (EC and IC) are numbered and indicated. Sequence alignments were manually edited (for details, see Methods). Of 14 motifs conserved in V1Rs (all of them single amino acids, identified by Rodriguez et al. 2002) eight are not conserved in ORs (cf. Niimura and Nei 2005) and consequently were chosen as analytical criterion here (*). (+) Residues conserved in ORs, some also in other GPCR families; (○) residues conserved in fish ora genes, but not in mammalian V1R genes.











