Evidence of transcription in biased regions
Click on table to view larger version.
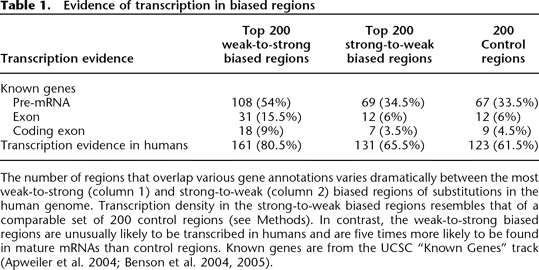
The number of regions that overlap various gene annotations varies dramatically between the most weak-to-strong (column 1) and strong-to-weak (column 2) biased regions of substitutions in the human genome. Transcription density in the strong-to-weak biased regions resembles that of a comparable set of 200 control regions (see Methods). In contrast, the weak-to-strong biased regions are unusually likely to be transcribed in humans and are five times more likely to be found in mature mRNAs than control regions. Known genes are from the UCSC “Known Genes” track (Apweiler et al. 2004; Benson et al. 2004, 2005).











