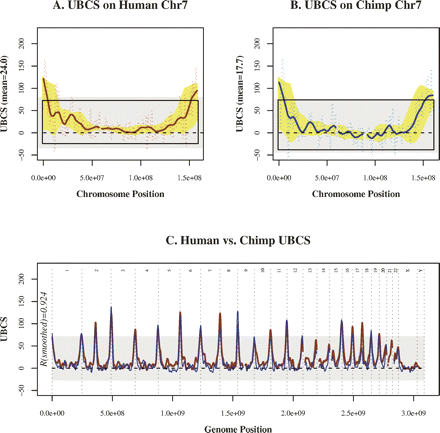
Patterns of substitution bias are nearly identical in human and chimp. (A) Unexpected biased clustered substitutions (UBCS) (faint line) for human chromosome 7 is above zero, indicating GC bias, along most (91.1%) of the chromosome and rises significantly at the distal ends. Smoothed UBCS (dark line) and 95% confidence band (yellow) are shown. The 95% confidence region is above the null expectation (zero) for more than half of the chromosome (61.8%). UBCS only exceeds the genome-wide 95% confidence interval (gray) near the telomeres. (B) Chimpanzee chromosome 7 has a remarkably similar profile (Spearman correlation ρ = 0.87). (C) The pattern of bias on chromosome 7 is mirrored on all autosomes (ordered sequentially; red, human; blue, chimp). Elevated UBCS near telomeres exceeds the human genome-wide 95% confidence interval (gray) on almost all autosomes. Here the chimpanzee sequence has been aligned to the human genome.











