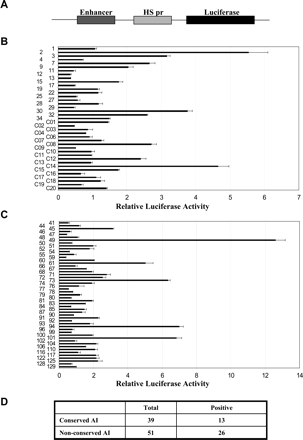
Enhancer activity assays of the conserved and nonconserved acetylation island sequences. (A) The construct used to test for enhancer activity. Each construct contains a 1.2-kb sequence encompassing the acetylation island sequence, which is cloned upstream of a luciferase reporter gene driven by a heat-shock gene promoter (HS pr). (B) Reporter gene activities of 39 acetylation island sequences that are conserved between human and pufferfish, following transient transfection of the constructs in Jurkat cells. The relative luciferase activity (x-axis) was determined by comparing the luciferase activity of the test construct to that of a similar construct without a putative enhancer insertion. The construct numbers (as shown in Supplemental Table S1) are indicated on the y-axis. Error bars indicate the standard deviation from the mean. (C) Reporter gene activities of 51 acetylation island sequences that are not conserved between human and mouse, as plotted in B. The construct numbers on the y-axis are described in Supplemental Table S2. (D) Summary of enhancer assay results. (AI) Acetylation island.











