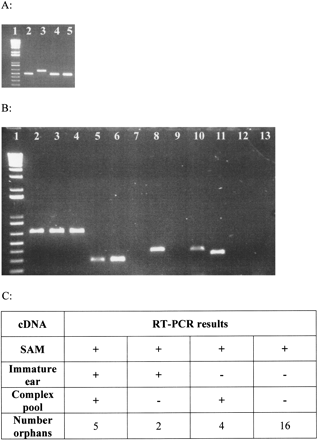
Experimental validation of the expression of orphan genes. (A) Test for genomic DNA contamination of cDNA. Primers that flank a 100-bp intron in the maize beta-tubulin6 (tub6) gene were used to amplify genomic DNA (lane 3), SAM cDNA (lane 2), immature ear cDNA (lane 4), and the complex cDNA pool (lane 5). (B) Examples of orphans with validated expression patterns. Primers designed to amplify (lanes 2–4) MAGI_80343, (lanes 5–7) MAGI_60450, (lanes 8–10) MAGI_75030, and (lanes 11–13) MAGI_30050 were used to amplify (lanes 2, 5, 8, 11) SAM cDNA, (lanes 3, 6, 8, 12) immature ear cDNA, and (lanes 4, 7, 9, 13) the pooled cDNA sample. (C) Summary of RT-PCR results for the 27 orphan genes. (+) Indicates that an RT-PCR product of the correct size was detected. Lane 1 of panels A and B contains the One KB Plus size standard (GIBCO BRL). Because primer dimers present in some lanes were cropped in both panels, the smallest size standard band shown is 200 bp.











