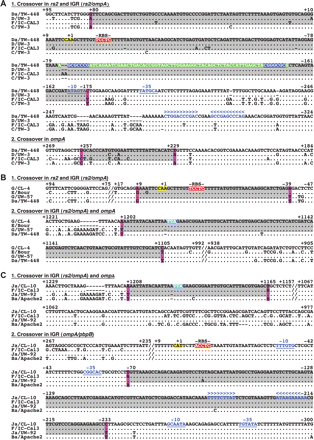
Nucleotide sequences of crossovers for reference strain Da/TW-448 (A) and clinical isolates G/CL-4 (B) and Ja/CL-10 (C). Numbers represent positions relative to the start codon of rs2 or ompA (highlighted in yellow). The stop codon of ompA is highlighted in blue. Crossover regions are highlighted in gray bordered by informative sites (fuchsia) from SimPlot/BootScan analysis. The tRNA is highlighted in green with inverted repeats (blue). Putative ribosomal binding sited (RBS) are highlighted in red. An imperfect and a perfect palindrome denote the putative terminators in blue below the symbols >>> <<<. Putative promotor regions are in blue characters and underlined; spacer A/T regions occur downstream from each −35 promoter element.











