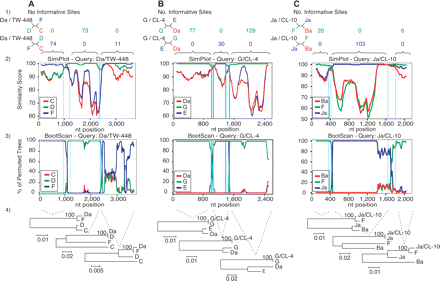
SimPlot representation of the putative crossover regions for the reference strain Da/TW-448 (A) and the clinical isolates G/CL-4 (B) and Ja/CL-10 (C). (1) Number of informative sites shared by the recombinant sequences (black) and the parental reference strains (blue and green). The outgroup sequence is shown in red. Four-member trees consistent with these sites are also shown for each region adjacent to the crossover region. (2) Similarity plot between each recombinant sequence and the respective parental reference strains, with a sliding window size of 200 bp and a step size of 20 bp. (3) BootScan analysis (window size 200 bp; step size 20 bp) showing the phylogenetic relatedness (% of permuted trees) between these sequences. For (2) and (3), the crossover regions are located between each pair of vertical lines, and nucleotides at the bottom of each plot correspond to alignment positions (not to chromosomal locations) of the continuous genomic regions analyzed. For Da/TW-448 and G/CL-4, the genomic region involved the rs2 gene to the ompA/pbpB IGR, while for Ja/CL-10, the region included only the rs2/ompA IGR, ompA, and the ompA/pbpB IGR. (4) Phylogenetic reconstructions for each specific region bounded by the recombination breakpoint region supporting each crossover (1000 bootstrapped trees).











