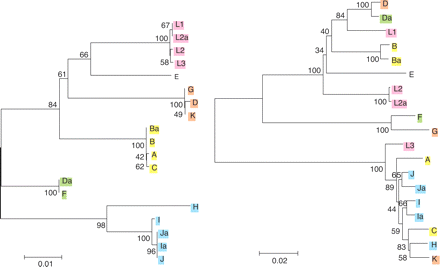
Phylogenetic reconstructions. Reconstructions of the evolutionary history of (left) CT049 and (right) ompA are based on the respective genomic sequences of C. trachomatis reference strains. Strains are color-coded based on the main lineages of the CT049 tree. Strains responsible for the three different disease groups of C. trachomatis (A, B, Ba, and C for trachoma; L1, L2, L2a, and L3 for invasive STDs; and H, I, J, Ja, I, and Ia for noninvasive STDs and likely persistent infection; Dean et al. 2000) form distinct clusters in the CT049 tree, but not in the ompA tree. They also cluster in trees for CT166 and pmpC (see Supplemental material), indicating the possibility of a clonal frame to which ompA does not belong (see Discussion). The values at the nodes are bootstrap that support levels representing the percentage of 1000 bootstrap resamplings in which the strains to the right form a clade. Amino acid trees (data not shown) are similar to the nucleotide trees.











