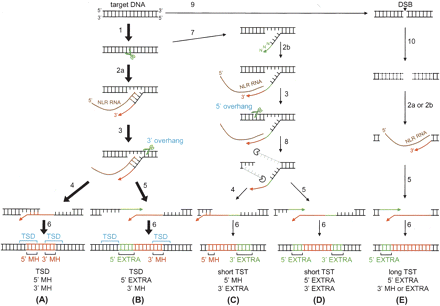
Possible pathways for NLR retrotransposition. Major pathways that result in TSD and 3′ MH stretches are indicated by bold arrows (A,B), and the others are indicated by thin arrows (C,D,E). Some other pathways, although not shown, are possible; for example, one that generates long TSTs and 5′ MH. The numbered arrows indicate the following reactions: (1) first-strand cleavage by NLR-encoded ENs, (2a) reverse transcription initiated with the help of annealing of target DNA and NLR RNA, (2b) reverse transcription primed by extra nucleotides at the 3′ end, (3) second-strand cleavage, (4) annealing of nascent NLR cDNA and target DNA, (5) addition of extra 5′ nucleotides, (6) sense-strand synthesis and ligation, (7) addition of extra 3′ nucleotides, (8) nucleolytic digestion of overhanging sequences, (9) introduction of a double-strand break (DSB), and (10) exonucleolytic digestion from the DNA ends. (EXTRA) Extra nucleotides added either the 5′ or 3′ end.











