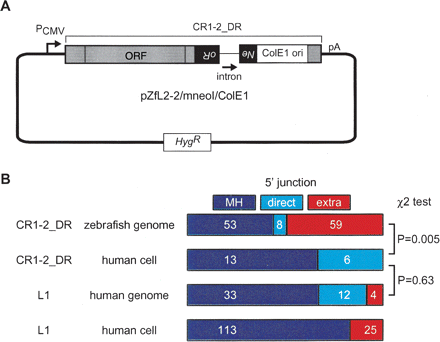
Retrotransposition assay in HeLa cells. (A) Construction of pZfL2–2/mneoI/ColE1. A full-length CR1–2_DR was placed under the control of the CMV promoter (PCMV) in the pCEP4 vector carrying the hygromycin-resistance gene (Hyg). The mneoI marker and ColE1 origin were inserted in the 3′ UTR of CR1–2_DR. The mneoI marker is a neomycin-resistant gene (Neo) interrupted by an insertion of an intron in the antisense orientation. This marker itself is in an antisense orientation relative to the NLR transcript. Thus, the vector does not confer the Neo phenotype, whereas CR1–2_DR retrotransposition, which includes splicing of the transcript, reverse transcription of that spliced RNA, and insertion of the synthesized cDNA into the genomic DNA, restores an intact Neo gene, converting the host cell to Neo. (B) Statistics for various NLR integrants. Integrants of CR1–2_DR in the zebrafish genome (top), those in HeLa cells using pZfL2–2/mneoI/ColE1 (second), L1 copies in transposons in the human genome (third), and de novo L1 insertions in human cultured cells (bottom) were categorized with regard to 5′ junctions. The frequency of extra 5′ nucleotides in de novo L1 insertions were reported previously (Symer et al. 2002; Gilbert et al. 2005). Because MH and direct joining were not distinguished in these reports, these integrants are represented as MH. The number of copies collected is indicated inside each rectangle. The P-values by χ2 tests for each pair are indicated at the right.











