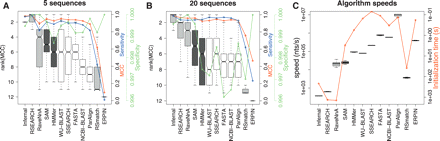
A comparison of the accuracy and efficiencies of homology search methods showing only the highest-ranking parameter settings for each algorithm from Supplemental Table 1. These were NCBI-BLAST (W7, 65%), WU-BLAST (W3), FASTA, ParAlign (65%), SSEARCH, HMMer (2.3.2, local), SAM (3.5, local), ERPIN, Infernal (0.7, local), RaveNnA, RSEARCH, and RSmatch. (A,B) Boxplots of algorithm ranks for the 5 and 20 sequence subsets, respectively. The blue curves show the median sensitivity, the green curve the median specificity, and the red curve the median MCC for each of the 12 programs. These accuracy values were computed by sampling either 5 or 20 sequences from the reference databases; these were used as input(s) to each algorithm for screening both the reference and a shuffled database. (C) Boxplots of algorithm speeds in nucleotides per second. The red curve shows median initialization times for the different programs. The single sequence, profile HMM, and RNA methods are displayed in unshaded, dark shaded, and lightly shaded boxes, respectively.











