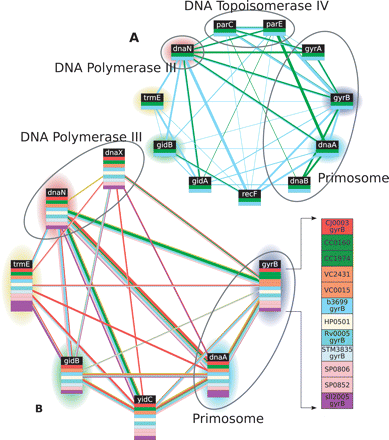
Two alignments of proteins involved in DNA replication. (A) A pairwise alignment between E. coli and C. crescentus includes several proteins involved in cell division as well as a conserved thiophene and furan oxidation protein. (B) A multiple alignment extends the pairwise alignment to include S. typhimurium, V. cholerae, C. jejuni, H. pylori, M. tuberculosis, S. pneumoniae, and Synechocystis. In this and subsequent figures, each colored box represents a protein and each vertical array of boxes represents an equivalence class; Græmlin hypothesizes that proteins in the same equivalence class performed the same function in the most recent common ancestor of the aligned species. To avoid clutter, individual proteins are not labeled, and, instead, each equivalence classis labeled with the consensus gene name of the proteins in it; as an example of the set of proteins aligned in an equivalence class, the detailed inset shows the specific proteins aligned to gyrB. Each protein is colored according to species, using the color code in Table 1; edges are also colored using the same scheme, and the width of each edge is proportional to its weight. In this figure, equivalence classes in the multiple alignment are highlighted the same color as the pairwise equivalence classes that they subsume.











