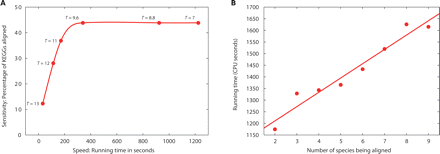
Running-time performance of Græmlin. (A) The speed sensitivity trade-off. Each point represents a run of Græmlin with d = 4 and different values of T. For each set of parameters, the x-axis plots the running time, and the y-axis plots the fraction of alignable KEGGs hit. (B) Progressive multiple alignment. Beginning with E. coli, we added species of increasing evolutionary distance to the multiple alignment. The pairwise running time is comparatively high because the two species aligned, E. coli and S. typhimurium, are the two most similar species and have many high-scoring alignments. In this manner, adding particularly close species to the alignment can lead to higher-than-average increases in running time, but over all species the average scaling will remain roughly linear.











