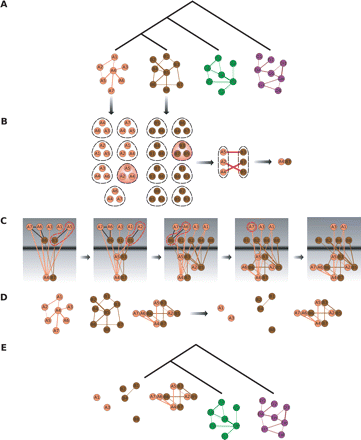
Outline of the Græmlin algorithm. (A) Shown here are four networks, together with their phylogenetic relationship. Græmlin will multiply align all four. (B) Græmlin first performs a pairwise alignment of the two closest species. It generates a set of d-clusters from each network; each node and its d − 1 closest neighbors constitute a d-cluster. Græmlin scores all pairs of d-clusters by finding for each pair the highest scoring mapping among nodes and selects the pairs that score greater than a user-specified threshold T; one particular high-scoring pair is highlighted together with the node mapping responsible for its score. Græmlin transforms all high-scoring pairs into seeds by aligning the two highest scoring nodes. (C) Græmlin extends the seed using a greedy algorithm; each extension step is shown in a gray box. At each step, Græmlin adds to the frontier all nodes connected to a node currently in the alignment; the frontier is shown in the upper half of each box. Græmlin adds to the alignment the pair of nodes or single node from the frontier that maximally increases the alignment score; the extension phase stops when no nodes from the frontier can augment the alignment score. (D) Græmlin transforms the resulting alignment, together with the unaligned nodes from the two original networks, into three generalized networks for use in the next phase of progressive alignment. (E) In the next step of progressive alignment, Græmlin will perform three pairwise alignments, one for each of the newly created generalized networks. Its running time will not scale by a factor of 3, however, as the total number of nodes in all networks remains roughly the same.











