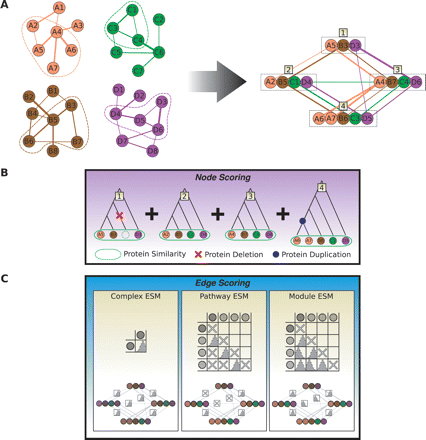
Method for scoring a multiple network alignment. (A) A sample multiple alignment. The four networks are from four different species. Each circle represents a protein, and edges link proteins that are hypothesized to interact; the width of an edge is proportional to the probability that an interaction is present. This sample alignment of the networks consists of four equivalence classes, numbered 1 through 4. (B) Node scoring method. Græmlin first scores each equivalence class independently by reconstructing the most parsimonious ancestral history of the involved proteins and then assessing penalties for sequence mutations and protein insertions, deletions, duplications, and divergences. Græmlin scores sequence mutations by using weighted sum-of-pairs scoring, obtaining pairwise scores based on BLAST results of the proteins; it scores all other events using heuristic constants. (C) Edge scoring method. Græmlin scores each edge using an Edge Scoring Matrix (ESM) as described in the text. For illustrative purposes, three alternative ESMs are shown, together with how Græmlin would score the alignment using each of them. The first ESM rewards protein complexes by specifying that edges between any pair of equivalence classes should have high weight; the matrix has only one cell because every edge is scored with the same distribution. The Complex ESM will score the alignment fairly highly but will penalize it because of the missing edges between equivalence classes 1 and 4 as well as 2 and 3. The Pathway ESM assigns a higher score to the alignment because it does not require high weight edges between all pairs of equivalence classes. It achieves this by a four-row matrix, where each label corresponds to a distinct node in a four-protein pathway. Edges between adjacent nodes in the pathway have high weights, and all other edges can have high or low weights without affecting the score; “don't care” distributions, symbolized by an X in the matrix, assign a score of 0 to those edges. The Module ESM assigns an even higher score to the alignment by conforming exactly to its structure; such an ESM is useful when a known module in one species is used as a query for searching another network.











