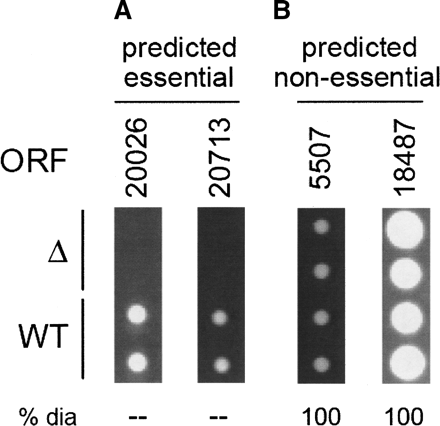
Observed colony sizes for knockout strains in S. mikatae, spotted in duplicate. ORFs were deleted with PCR and transformed into diploids. Sporulation was induced and tetrads dissected to recover haploid cells carrying the deletion of interest. Each column represents a different deletion strain; the top two spots represent colonies grown from deletion strains, while the bottom two spots are wild-type colonies. Four ORFs were selected for knockout mutagenesis: (A) two high-scoring ORFs representing our top essentiality predictions; deletions of predicted essential ORFs were non-viable; (B) two low-scoring ORFs likely to represent non-essential genes; deletions of predicted non-essential ORFs displayed wild-type growth. (ORF) S. mikatae ORF deleted in this strain. (Δ) Growth of knockout strain (deletion of ORF listed above). (WT) Growth of wild-type strain. (% dia) Percent of mutant colony diameter compared with WT for those with slow growth.











