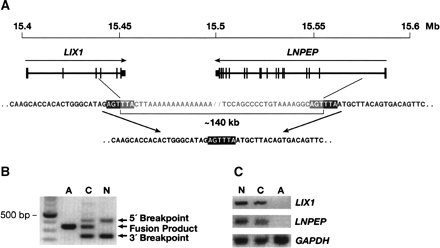
An ~140-kb deletion abrogates LIX1 and LNPEP expression in SMA affected cats. (A) A schematic of the genomic organization of LIX1 and LNPEP on FCA A1q. Arrows indicate the direction of transcription of each gene. The scale of chromosome coordinates is from the region of conserved synteny in the dog genome (CFA 3) (UCSC Genome Browser; July 2004 assembly). Sequence of the deleted and normal alleles are aligned showing the precise breakpoints below the schematic. The immediately flanking 6-bp sequence, AGTTTA, is in bold. (Note that the genome browser did not itself identify exon 1 of LNPEP by homology with RefSeq genes, but limited homology was identified by BLAT search of the dog genome with a cat sequence that was 82% identical over 300 bp of the human exon 1 and flanking 5′-UTR and intron 1 sequences). (B) PCR products produced from genomic DNA in a multiplex reaction designed to amplify across both deletion breakpoints when the normal allele (lane N) was present. A third product corresponding to the affected cat sequence shown in A amplified when the deleted allele was present, either in homozygous affected (lane A) or heterozygous carrier (lane C) cats. (C) RT–PCR products amplified from cervical spinal cord ventral horn RNA of a genetically normal cat (lane N), a clinically normal carrier (lane C), and an SMA affected cat (lane A). PCR primers for LIX1 were from exons 1 and 4, LNPEP primers were from exons 14 and 15, and GAPDH primers were from exons 6 and 8.











