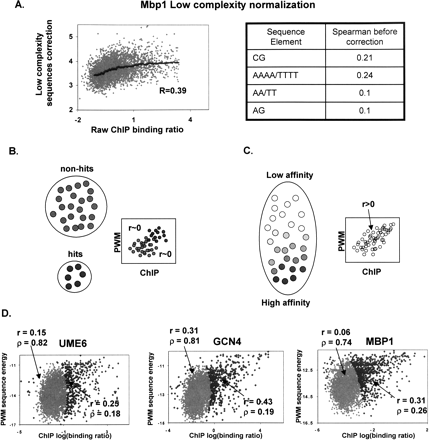
Testing the digital model. (A) Normalizing ChIP data. PREGO performs internal normalization of the ChIP data to eliminate any correlation of the binding ratios to single or dinucleotide composition or to low complexity sequences [typically poly(A) or poly(T) tracts]. Shown are the scatter and trend of the raw Mbp1 ChIP binding ratio versus the inferred correction, involving contribution from several dinucleotides and an AAAA/TTTT motif. The Spearman correlation of each of the sequence features used in the normalization and the ChIP data is also shown (right). (B,C) Discrete versus analog models. If TF–gene interactions can be reasonably approximated as either occurring or not occurring (hits or non-hits), then the joint distribution of ChIP and PWM predictions should reflect zero covariance inside such two ideal subsets of the genome (left). If ChIP and PWM provide quantitative estimations on in vivo binding affinity, then no partition of the genome can eliminate their correlation (right). It is therefore possible to test the validity of the digital assumption by fitting two distributions to the data and analyzing their parameters. (D) ChIP-sequence correlation reflects an analog behavior. Analysis of the ChIP/PWM joint distributions for three TFs reveals that their quantitative correlation cannot be explained as a consequence of the mixture of two distributions (Methods). Shown are inferred maximum likelihood distributions for hits (darker) and non-hits (brighter). The mixture coefficients (ρ) and correlation coefficients (r) are indicated. The analysis suggests that about one-fifth of the genome is influenced by each of the TFs, and that for at least one-fifth of the genome, ChIP- and sequence-based estimations of affinity are correlated in a quantitative fashion.











