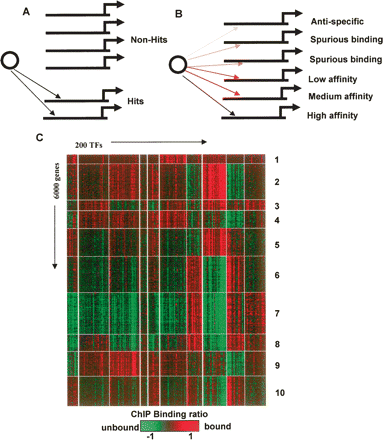
The transcriptional program in yeast: digital or analog? According to the prevalent “digital” hypothesis for transcriptional regulation (A), complex regulatory programs are described using wiring diagrams that associate TFs to genes deterministically. In the alternative “analog” model (B), many TFs may affect each gene at drastically different levels of specificity. Two-way clustering of 200 ChIP binding profiles and 6000 yeast genes (C) reveals groups of genes with remarkably similar binding ratios in all 200 ChIP experiments. Few of the entries in the homogeneous submatrices represent high-specificity TF–gene associations. The clusters and their association with biological functions (Supplemental Table 1) suggest that ChIP experiments may reflect complex and functionally meaningful organization of low-affinity TF–gene interactions.











