Regions of mouse genome represented on microarray
Click on table to view larger version.
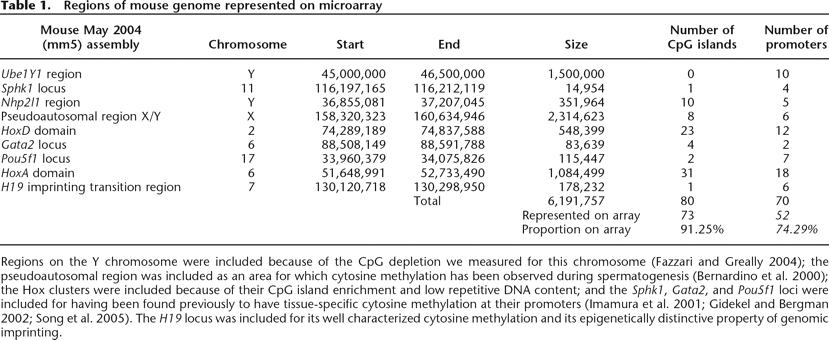
Regions on the Y chromosome were included because of the CpG depletion we measured for this chromosome (Fazzari and Greally 2004); the pseudoautosomal region was included as an area for which cytosine methylation has been observed during spermatogenesis (Bernardino et al. 2000); the Hox clusters were included because of their CpG island enrichment and low repetitive DNA content; and the Sphk1, Gata2, and Pou5f1 loci were included for having been found previously to have tissue-specific cytosine methylation at their promoters (Imamura et al. 2001; Gidekel and Bergman 2002; Song et al. 2005). The H19 locus was included for its well characterized cytosine methylation and its epigenetically distinctive property of genomic imprinting.











