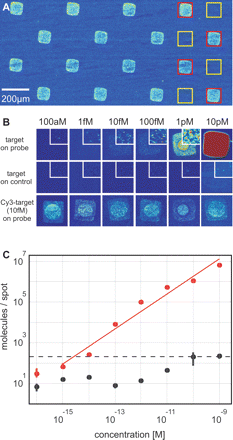
Validation of the system. (A) 60mer probe (red squares) and target (yellow squares) oligonucleotides were spotted. Specific signals after hybridization with 100 fM Cy5-labeled target oligonucleotide show a spot-to-spot variation of only 13%. For illustration, data were software-binned (10 × 10). (B) Fluorescence images of spots hybridized with different concentrations of Cy5-labeled target-oligonucleotide. Individual Cy5-labeled oligonucleotide molecules can be detected in the range of from 100 aM to10 fM; at higher concentrations, single molecule signals begin to overlap. (Insets) Details within the spots at 3.5-fold higher magnification. In the second row, the corresponding images of control spots are shown. A constant concentration of Cy3-labeled target-oligonucleotide was included in all experiments (third row). (see Supplemental Fig. S2 for images of higher target concentrations). (C) Number of specifically hybridized (red) and unspecifically adsorbed (black) oligonucleotide molecules per spot vs. sample concentration. Three independent experiments, each containing 16 replicate spots, were performed for each concentration. The surface density increases linearly with c over six orders of magnitude (surface-density ∝ c0.86). Assuming a minimum signal-to-noise ratio of 10 as criterion for reliable analysis (dashed line), a detection-limit of c = 1.3 fM can be specified. Error bars indicate the standard deviations of the signals. Note that due to the logarithmic scaling, the error bars are only visible at low concentrations (see also Supplemental Table S1).











