Summary of fractionation bias by α-region
Click on table to view larger version.
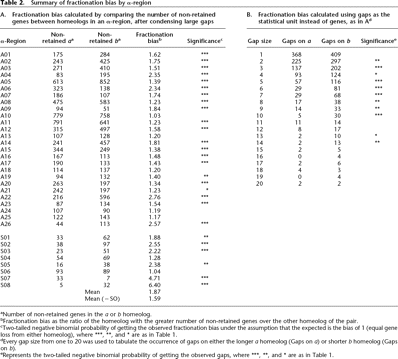
aNumber of non-retained genes in the a or b homeolog.
bFractionation bias as the ratio of the homeolog with the greater number of non-retained genes over the other homeolog of the pair.
cTwo-tailed negative binomial probability of getting the observed fractionation bias under the assumption that the expected is the bias of 1 (equal gene loss from either homeolog), where ***, **, and * are as in Table 1.
dEvery gap size from one to 20 was used to tabulate the occurrence of gaps on either the longer a homeolog (Gaps on a) or shorter b homeolog (Gaps on b).
eRepresents the two-tailed negative binomial probability of getting the observed gaps, where ***, **, and * are as in Table 1.











