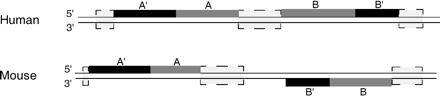
Pair generation. Example of the generation of two pairs, pair A and pair B. The black boxes are matching alignments, the light gray boxes are repeats, gaps, or alignments, and the dark gray boxes are the pairs, A and B. Pair A is downstream of the A′ +/+ alignment, and both regions end at a repeat, gap, or another alignment. Pair B is upstream of the B′ +/− alignment; therefore, both regions are placed at the 5′-end of the alignment; again, the length of these pairs is limited by repeats, gaps, or another alignment. The remaining nonboxed lines are nonconserved regions, which are not adjacent to matching alignments.











